
[ad_1]
A Ct worth, often known as cycle threshold, is essential in quantitative PCR. It represents the cycle quantity at which a pattern’s response crosses a fluorescence threshold, indicating the detection of the goal nucleic acid. Decrease Ct values point out larger goal sequence quantities, whereas larger Ct values counsel decrease quantities or points. Varied components can have an effect on Ct values, however normalization strategies like Delta-Delta Cq assist differentiate biological-relevant modifications from technical points by evaluating pattern Ct values to reference genes.
If you’re performing quantitative PCR (qPCR), then it’s essential you perceive what a Ct worth is. We’ll take you thru its important position in qPCR, its various names, the way it’s calculated, and what it means when all of it appears to go a bit fallacious.
The Many Names of the Ct Worth
Earlier than we dive into explaining what a Ct worth is, we wish to take a second to spotlight that that worth has been given a number of names over time, together with:
- Ct – cycle threshold/threshold cycle
- Cp – crossing level
- TOP – take-off level
- Cq – quantification cycle
These values are all the identical, simply with totally different names. To standardize qPCR nomenclature, the MIQE (minimum information for publication of quantitative real-time PCR experiments) pointers advocate utilizing Cq worth. These pointers standardize experimentation and reporting, which is important provided that quantitative real-time PCR is used to investigate medical samples and diagnose gentle and extreme illness, notably SARS-CoV-2 infections. Due to this fact, all through this text, we’ll use Cq solely. [1]
What’s the Cq Worth?
Actual-time PCR (typically referred to as qPCR) is often carried out to quantify absolutely the quantity of a goal sequence or to check relative quantities of a goal sequence between samples. This method displays the amplification of the goal in actual time through a target-specific fluorescent sign emitted throughout amplification.
Regardless that real-time PCR fluorescent dyes and probes must be sequence-specific, a substantial quantity of background fluorescence happens throughout most real-time PCR experiments. It’s important to bypass or account for this background sign to glean significant details about your goal. This problem is addressed by two values in real-time PCR: the edge line and the Cq worth.
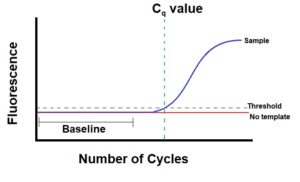
- The edge line is the extent of detection or the purpose at which a response reaches a fluorescent depth above background ranges. Earlier than conducting PCR, you (or the software program in your cycler) set a threshold degree. That is actually a line in your graph that represents a degree above background fluorescence, that additionally intersects your response curve someplace originally of its exponential section (Determine 1).
- The Cq worth is the PCR cycle quantity at which your pattern’s response curve intersects the edge line. This worth tells what number of cycles it took to detect an actual sign out of your samples. Actual-time PCR runs could have a response curve for every pattern and due to this fact many Cq values. Your cycler’s software program calculates and charts the Cq worth for every of your samples.
Cq values are inverse to the quantity of goal nucleic acid in your pattern and correlate to the variety of goal copies in your pattern. Decrease Cq values (usually under 29 cycles) point out excessive quantities of the goal sequence. Larger Cq values (above 38 cycles) imply decrease quantities of your goal nucleic acid. Excessive Cq values may also point out issues with the goal or the PCR set-up, as outlined later within the pitfalls part of this text.
Your PCR instrument will accumulate fluorescence knowledge throughout every cycle. After about 15 cycles, you’ll have a good suggestion of your background fluorescence degree – this can seem as a horizontal line ranging from the zero cycle level. The edge degree might be simply above this however on the level the place your samples begin transferring into the exponential section of PCR amplification. Immediately, laptop software program calculates this actual level, and all fashionable real-time cyclers have an computerized threshold line setting.
Actual-time PCR information the quantity of fluorescence emitted through the response the place all PCR elements are considerable. On this means, Cq values are often constant throughout replicates in real-time PCR. By the point the PCR response endpoint is reached, collected inhibitors, inactivated polymerases, and limiting reagents create loads of variation in endpoint values, and this is the reason standard PCR can’t be used quantitatively.
Components That Can Affect Ct Values
Many components can have an effect on your Cq values. Some variations in Cq values between your samples might be because of organic occasions, e.g., up/down-regulation of your goal gene in response to a therapy. Nonetheless, Cq values are simply as simply influenced by the preparation of the PCR response and the PCR elements themselves. The most typical pitfall areas are:
1. Grasp Mixes
Fluorescence emission might be affected by pH and salt focus in an answer. Any change in fluorescence emission will naturally change your Cq values. Due to this fact, be sure you solely use high-quality PCR elements, and if utilizing selfmade options, verify the pH and monitor salt precipitation earlier than every experiment.
2. Passive Reference Dyes
Response values are the ratio of the fluorescence of your FAM (reporter) dye to your ROX (passive reference) dye. Decrease quantities of ROX produce larger response values, assuming FAM fluorescence doesn’t change.
3. Response Effectivity
PCR response effectivity depends on the grasp combine efficiency, the specificity of the primers, the primer annealing temperature, and the pattern high quality. Normally, PCR effectivity above 90% is appropriate. PCR effectivity of 100% signifies that the goal sequence of curiosity doubles throughout every cycle. Good PCR effectivity coincides with a change of three.3 cycles between 10-fold dilutions of your template.
To find out the PCR effectivity for every primer pair, run serial dilutions of your template with 5 10-fold dilution steps and calculate the R2, which is a statistical measure that describes how nicely one worth can predict one other. For PCR effectivity near 100%, your R2 worth must be better than 0.99.
Run at the very least three replicates for every level in your normal curves. A better replicate quantity is very necessary for low copy quantity enter, the place variations throughout replicates are extra possible.
4. Different Points
Assuming you may have dominated out the three components talked about above, the most typical causes of late Cq values are:
- Too little template – attempt utilizing extra template.
- Suboptimal nucleic acid isolation – think about your nucleic acid isolation protocol, quantify your DNA, run an agarose gel, or attempt one other protocol/equipment.
- Poor reverse transcriptase exercise throughout cDNA synthesis – reverse transcriptase is delicate to degradation. Order a brand new one.
- RNA/cDNA degradation – preserve your workspace clear, enhance your RNA dealing with conduct, and keep away from a number of freeze-thaw cycles of cDNA.
- PCR inhibition. [2]
Extra causes embody an infection or contamination in your cell line/tradition, however these points are often noticed forward of nucleic acid isolation.
Calling Delta-Delta Cq
To make sure that the variations in Cq values are because of actual organic modifications and never technical points, you will have to normalize your outcomes. The preferred normalization methodology is called “Delta-Delta Ct” or the Livak methodology. Right here, you examine the Cq values of your pattern to the Cq values of a number of reference (housekeeping) genes.
It’s crucial to select reference genes whose expression ranges will not be anticipated to alter throughout your experiment. Widespread housekeeping genes embody actin, alpha-tubulin, GAPDH, and ubiquitin. It’s smart to make use of at the very least two reference genes and keep in mind that what could also be a reference gene for one examine is probably not appropriate for an additional.
The Delta-Delta Cq methodology makes a key assumption—that the amplification (PCR) efficiencies of your reference and goal samples are nearly 100% and inside 5% of one another. Different normalization strategies embody the Delta-Cq methodology and the Pfaffl Methodology. You may learn extra concerning the qPCR knowledge evaluation strategies right here.
Ct Values Summed Up
Hopefully, this text has helped you get a grip on Cq values. If you’d like extra info on qPCR, be sure you take a look at our high 11 qPCR Papers Each Researcher Ought to Know.
For a extra complete information to PCR and PCR parameters, obtain our free PCR fundamentals eBook and turn into an professional.
Initially printed in July 2015. Up to date and republished in November 2020. Revised and up to date once more in October 2023.
References
- Bustin SA, et al. The MIQE pointers: minimal info for publication of quantitative real-time PCR experiments. Clin Chem. 2009. 55(4):611–22. doi: 10.1373/clinchem.2008.112797.
- Schrader C, et al. PCR inhibitors – incidence, properties, and elimination. J Utilized Microbiology. 113(5):1014–26. doi: 10.1111/j.1365-2672.2012.05384.x
[ad_2]